Welcome...

Announcements will be made on this site and all resources, e.g. papers for the seminar, code that is being used in the lectures, lecture notes and slides will be made available here for download in the respective sections.
If you have further questions contact me.
What is this course about?

Topics include:
The course will cover various topics in complex systems research for example:
- self-organization
- pattern formation
- critical phenomena
- growth processes
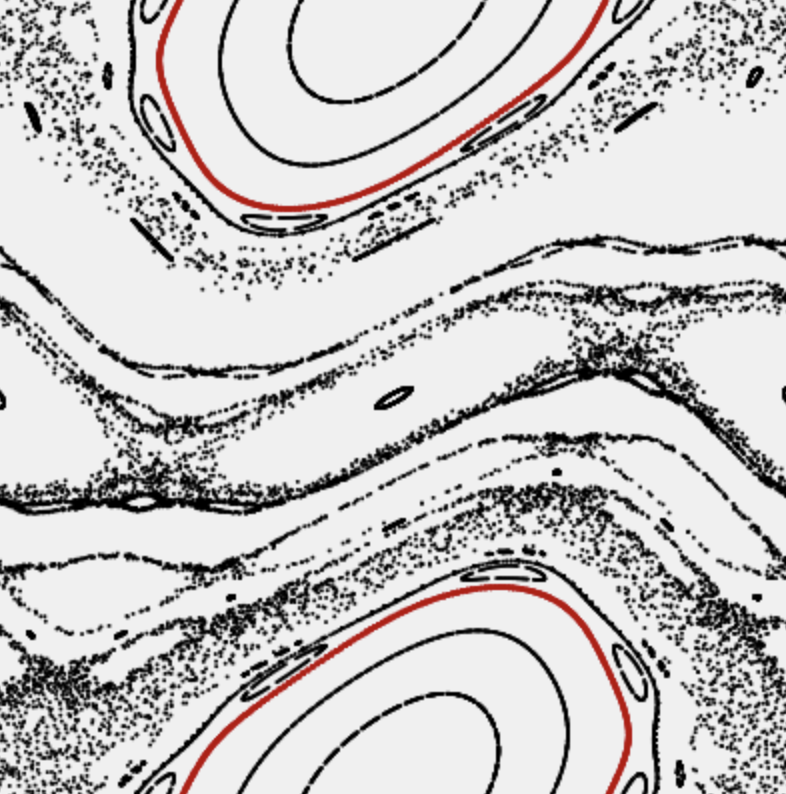
- deterministic chaos
- synchronization
- collective behavior
- biological networks
- self-similarity
General Information
The module consists of the Lecture Introduction to Complex Systems, the Seminar Complex Systems in Biology and the Practical Course Computer Simulations and Modeling of Complex Systems. Information on each of these is available in the respective sections.
The module will provide an interdisciplinary perspective on complex dynamical phenomena in biological systems. A solid theoretical/modeling foundation will be provided as well as a number of applications to biological systems.
The module Complex Systems in Biology is part of the Masters Biophysics program and can count as a mandatory module (Pflichtmodul) in theoretical biophysics II or alternatively as an elective module (Wahlpflichtmodul) for the program.
Although the lecture notes and this website is maintained in English the module will be taught in German.
The Lecture

The lecture will also introduce all applied topics covered in the module and discuss their properties and dynamic aspects.
The Seminar
The seminar is designed for students to focus more deeply and individually on a specific research project from the range of topics discussed in the lecture. Each student will pick a Complexity Explorable and study the system it describes and related papers in more detail.
Each student will have to give a presentation of the explorable. Because of Covid-19 the presentations cannot be given in class. Each student can make a recording of the presentation and turn it in. The presentations will be collected on the Seminar page.
The Lab Course

Requirements
The basic requirements for the successful completion of the course is a basic knowledge of calculus and very basic knowledge of differential equations. Programming skills are not required but helpful.
If you are uncertain whether you meet the requirements, contact me.
Every student requires a laptop computer. Most students have a laptop, for those of you who do not have a laptop I can provide one.
Contact
The best way to contact me is by Email complexsystems2021@gmail.com.